‘Embrace the Ditch,’ and Other Lessons Learned in Duke CEE’s Overture Engineering
Civil and environmental engineering students learn to design buildings within less-than-optimal parameters in a collaborative capstone course
We’re sorry, but that page was not found or has been archived. Please check the spelling of the page address or use the site search.
Still can’t find what you’re looking for? Contact our web team »
Read stories of how we’re teaching students to develop resilience, or check out all our recent news.

Civil and environmental engineering students learn to design buildings within less-than-optimal parameters in a collaborative capstone course

On a Star Wars-themed field of play, student teams deployed small robots they had constructed
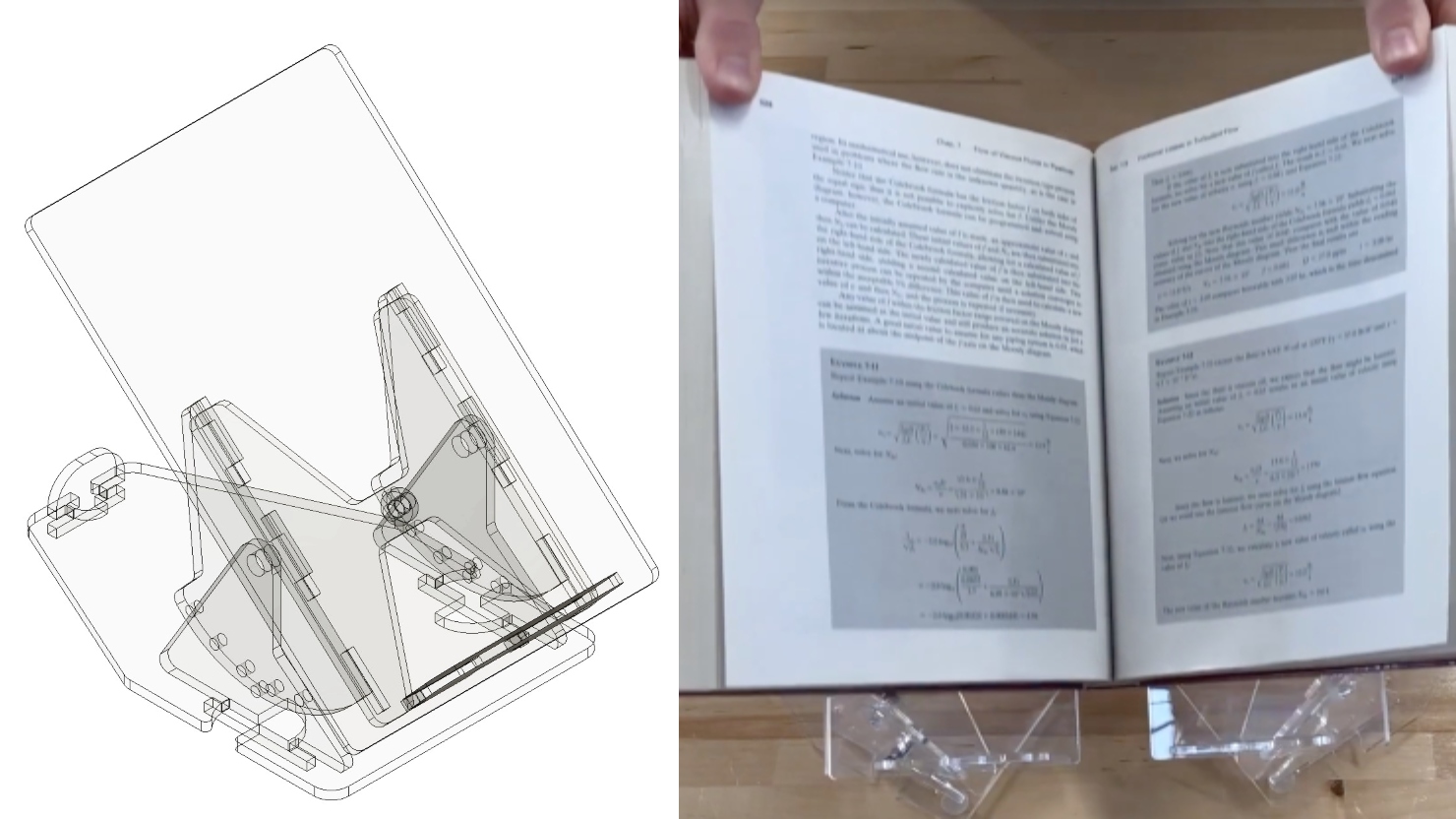

Two projects from First-Year Design course are patent-pending. Student surveys suggest the course also fosters teamwork, leadership and communication skills.
Apr 20
ABOUT THIS EVENT Drop by the 1st floor MakerLab at the Durham County Main Library on Saturday, April 20th anytime between 10am – 2pm to explore nanotechnology and pick up […]
10:00 am – 10:00 am
Apr 22
Learn what goes on behind the scenes when you submit your manuscript!Tips on submitting to the right journal, discuss peer review management, making editorial decisions, insight for communicating with editors […]
10:30 am – 10:30 am Fitzpatrick Center Schiciano Auditorium Side A, room 1464
Apr 22
The healthcare sector stands at a pivotal crossroads, reminiscent of Charles Dickens’ portrayal of contrasts in “A Tale of Two Cities”. Today’s healthcare landscape is marked by stark dichotomies: we […]
12:00 pm – 12:00 pm Teer 203